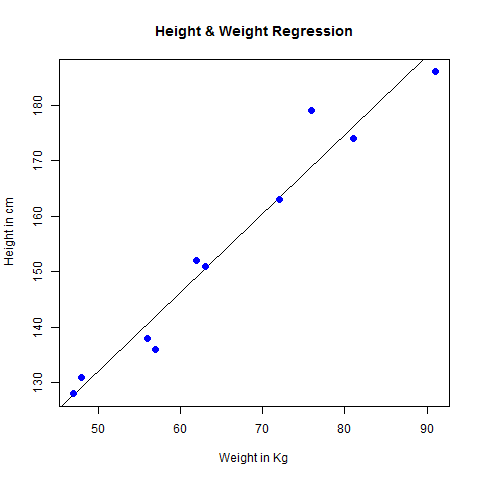
This step is performed using the FindNeighbors() function, and takes as input the previously defined dimensionality of the dataset (first 10 PCs). Briefly, these methods embed cells in a graph structure - for example a K-nearest neighbor (KNN) graph, with edges drawn between cells with similar feature expression patterns, and then attempt to partition this graph into highly interconnected ‘quasi-cliques’ or ‘communities’.Īs in PhenoGraph, we first construct a KNN graph based on the euclidean distance in PCA space, and refine the edge weights between any two cells based on the shared overlap in their local neighborhoods (Jaccard similarity). Our approach was heavily inspired by recent manuscripts which applied graph-based clustering approaches to scRNA-seq data and CyTOF data. However, our approach to partitioning the cellular distance matrix into clusters has dramatically improved. Importantly, the distance metric which drives the clustering analysis (based on previously identified PCs) remains the same. Seurat v3 applies a graph-based clustering approach, building upon initial strategies in ( Macosko et al). For example, performing downstream analyses with only 5 PCs does significantly and adversely affect results.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
